Excited to share that our paper "Many Needles in a Haystack: Active Hit Discovery for Perturbation Experiments" has been accepted at ICML 2026 — see you in Seoul! 🇰🇷
CRISPR and single-cell perturbation screens can test thousands of genes, but budgets are tight and the genes that actually move a phenotype are rare and scattered. The hard question is no longer can we screen? — it's which perturbations should we run next?
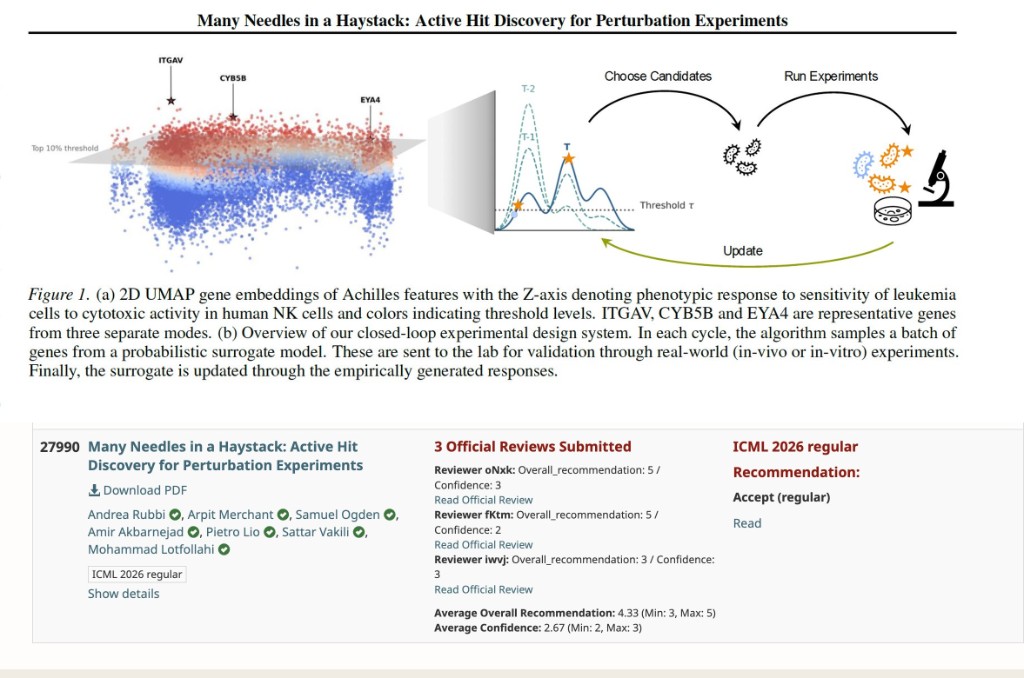
We tackle this with a lab-in-the-loop design loop: an AI model proposes a batch of genes to perturb, the lab runs them, the readouts update the model, and the cycle repeats — each round getting smarter about where the hits are hiding.
The catch is that standard active-learning methods chase a single "best" gene, but biology rarely works that way: hits sit in multiple pathways and cellular contexts. Our method, Probability-of-Hit, instead asks the right question — which genes are most likely to cross the hit threshold? — and recovers more biologically meaningful hits across five real immunology screens (T-cell activation, NK cytotoxicity, tau, SARS-CoV-2).
The upshot for experimentalists: more hits per plate, fewer wasted wells.
Congratulations to first authors Andrea Rubbi and Arpit Merchant, postdoc Sam Ogden, Amir Hossein Hosseini Akbarnejad, and thanks to Pietro Liò and Sattar Vakili for the great collaboration.
