Mixture-of-Experts (MoE) is a powerful way to scale large language models (LLMs): instead of running the full model for every token, a router activates only a few "experts," giving more capacity at roughly the same compute.
But routing is still a sore spot. Most MoE systems use Top-k + Softmax, where expert selection is discrete—so you don't get clean end-to-end gradients. In practice, this can lead to unstable routing, calibration issues, and uneven expert usage.
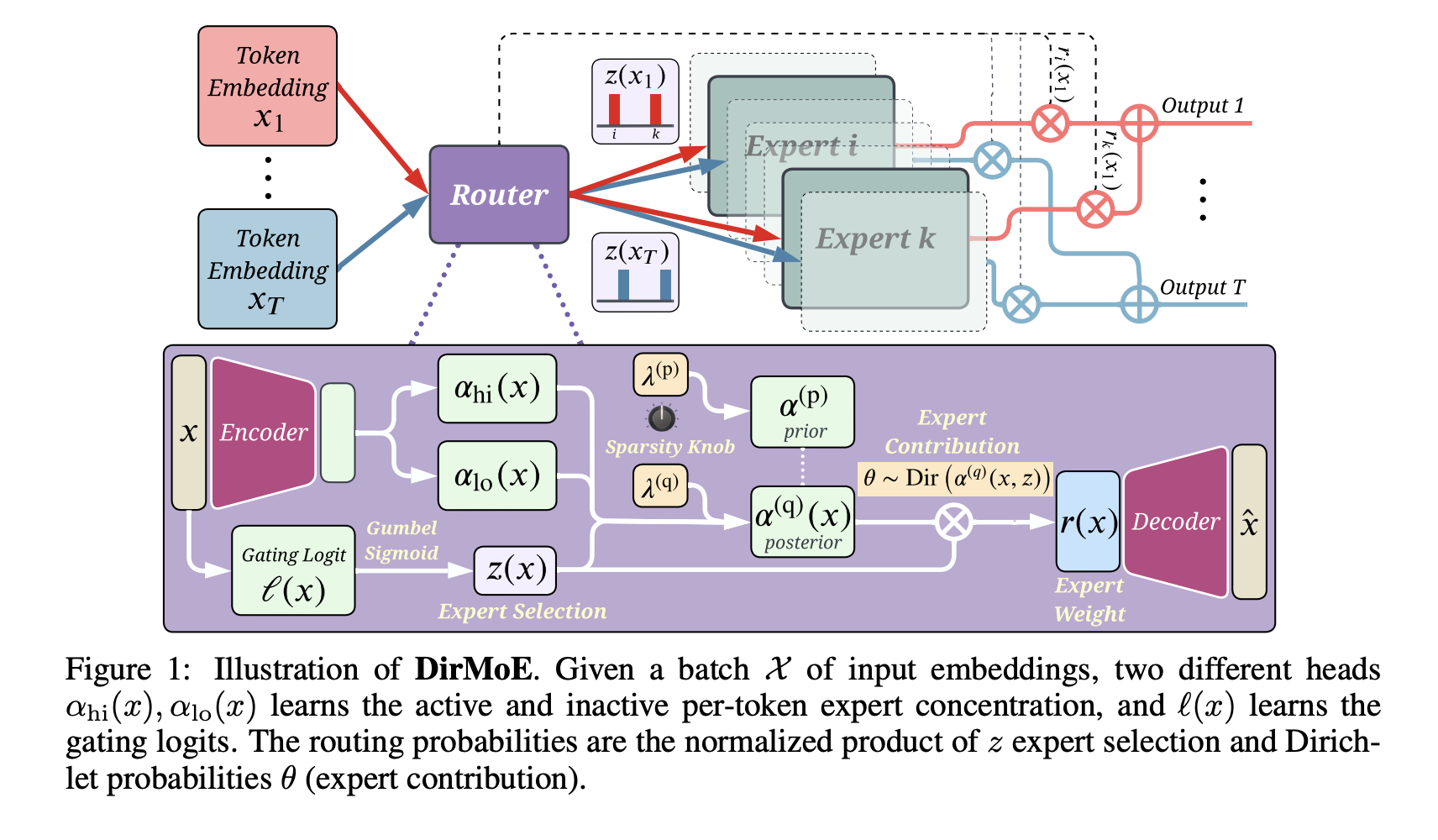
In our ICLR 2026 paper, we introduce DirMoE — a fully differentiable probabilistic router that separates which experts fire (Bernoulli) from how their weights are assigned (Dirichlet). We also add a simple "sparsity knob" (Simpson-index penalty) to control the expected number of active experts, without relying on load-balancing losses that can homogenize experts.
Results: DirMoE matches or exceeds vanilla MoE throughput (no extra bottlenecks), is strong/competitive on zero-shot benchmarks (ARC, BoolQ, PIQA, …), and leads to clearer expert specialization (interpretable domain focus like ArXiv/Books/GitHub code).
Led by Hesam Asadollahzadeh and Amirhossein Vahidi.

